1. Introduction
The fabrication of nanoparticles has a tremendous importance in various research fields of nanoscience and nanotechnology. Wherein, the synthesis of metallic copper and copper oxide particles are considered to be of increasing interest due to their high electrical, catalytic, sensing and surface properties as well as high antibacterial and antifungal activities, low toxicity, chemical stability1-4) and etc. For synthesis purposes, there are various methods used to produce nanoparticles, such as, co-precipitation, chemical reduction, electrochemical reduction, laser ablation, mechanical milling, radiation reduction, thermal decomposition and etc.5-10) Among these processes, the chemical reduction is usually considered to be ideal for its costeffectiveness and easy processing to get a narrow size distribution of nanoparticles by following the optimum process conditions, such as by varying the molar ratios of reducing agent and stabilizer with the precursor.11) The Cu nanoparticles produced at standard pressure without an inert atmosphere are easily oxidized and aggregated into copper oxides which are considered to be of more thermodynamic stability as compared to pure copper. To avoid this situation, they are produced using starch as a capping agent which acts to protect them from oxidation and aggregation.12) The use of starch as a stabilizer is measured to be beneficial for its low cost, high solubility in water, high capability to form complexes with metal ions and have a large number of hydroxyl functionalities and weak interaction with the metal surfaces to protect them at high temperature.
In addition, as compared to other methods, the chemical reduction method is considered to be promising wet chemistry synthesis approaches to produce metal nanoparticles with the ability to provide a favorable atmosphere for controlling the chemical reaction. In such a way, the production of nanoparticles along with a narrow size distribution can be achieved under easily controlled reaction rate.13)
The present work also deals with the preparation of CuNPs by a simple chemical method using copper salt (precursor), ascorbic acid (reducing agent) and starch (stabilizer). The aim of this study is focused on optimizing the process conditions, antibacterial and antifungal activities. The detailed analysis of the CuNPs using XRD, SEM and FTIR is also performed.
2. Experimental Procedure
The chemicals purchased from Sigma-Aldrich such as copper salt (CuSO4.5H2O), ascorbic acid (C6H8O6), Starch (C6H10O5) and caustic soda (NaOH) as of analytical grade. An aqueous solution was prepared in deionized water.
For preparation of CuNPs, the procedure of chemical method was followed.5) Wherein, a synthesis solution was prepared by mixing 50 mL of 0.1 M CuSO4. 5H2O solution into 60 mL of 1 % (0.6 g) starch solution and stirred for a time interval of 30 min as shown in Fig. 1(a, b). Then 25 mL of 0.2 M ascorbic acid solution was put into the same solution under constant stirring. During the mixing process, the color of the mixture solution changed from yellow to white as shown in Fig. 1(c). Finally, 25 mL of 1 M NaOH was gradually added to the synthesis solution under continuous magnetic stirring and heating at about 80 °C using hot plate for time duration of 2 hrs. Due to the addition of NaOH solution, the color of the synthesis solution changed to ocher (dark red) color as shown in Fig. 1(d). After completion of the reduction process, the solution was removed from the hot plate and allowed to cool and settle down CuNPs at room temperature for overnight. The supernatant solution was removed carefully and the precipitate was filtered and washed thoroughly with deionized water and then repeatedly with ethanol to get ride of starch present in the nanoparticles. The resulted product was dried at room temperature and then was put into a glass vial and was stored in a desiccator for further analysis as shown in Fig. 1(e).
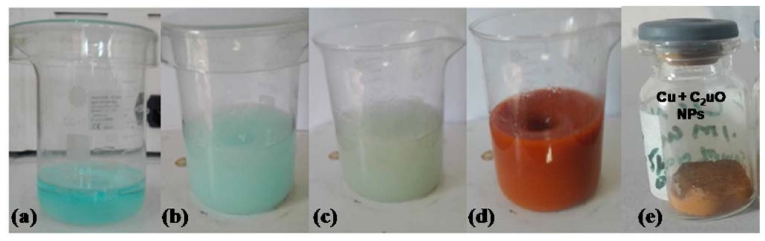

Fig. 1
(a) 0.1 M CuSO4.5H2O in 50 ml deionized water, (b) addition of 1 % Starch solution in 60 ml deionized water into a, (c) addition of 0.2 M ascorbic acid in 25 ml deionized water into b and (d) 1 M NaOH solution in 15 ml deionized water into c synthesis solution and (e) the powdered CuNPs.
The CuNPs in powder form were used in the X-ray diffraction analysis. The XRD pattern of the sample was obtained by a diffractometer (JEOL X-ray diffractometer, model JDX-3532, Japan) using X-rays; CuKa (λ=1.5418 Å). The crystallite size was found out by the following Scherrer equation (used for measuring nanoparticles sizes);
Where K is known as Scherrer constant or shape factor (value of about 0.9), λ for wavelength (0.15418 nm), β for line broadening at half the maximum intensity (FWHM) in radians, θ for Bragg angle, (the position of maximum diffraction peak) and d for the mean size of crystallite in nanometers. The surface morphology of the CuNPs was examined under a Scanning Electron Microscope (SEM, J-5910, JEOL). Infrared Spectroscopy (ALPHA Eco-ATR FTIR spectrometer, Bruker) was used to characterize the functional groups of the resulted CuNPs in powder form.
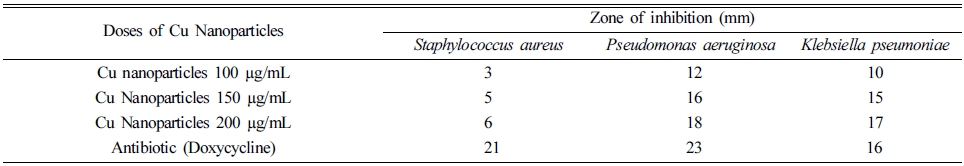
The synthesized CuNPs were tested for antibacterial activity against pathogenic bacterial strains both Gramnegative (Klebsiella Pneumoniae and Pseudomonas aeruginosa) and Gram-positive (Staphylococcus aureus) by using the agar well diffusion method.14) The agar molten plates with 24 hrs mature broth culture of individual bacterial strains were taken to make bacterial lawn carefully. Different concentrations of CuNPs such as 100 μg/100 μL, 500 μg/150 μL, and 1000 μg/200 μL were added to each of well containing a plate of bacterial strain to assess the activity of the CuNPs. The standard antibiotics Doxycycline was added to the Dimethyle sulfoxide (1 mg/1 mL) of concentration 50 μL to the center well plate for positive control against the bacterial pathogens. Then the plates were incubated at 37 °C for 24-36 hrs. The zone of inhibition was measured in millimeters (mm) of the every well. Triplicates were maintained in every concentration and the mean values were also calculated for the ultimate antibacterial activity.
For the growth media of fungi, 3.25 g SDA (Sabrous dextrose agar) was added in 50 ml distilled water and autoclaved for 15 min at 121 °C and at 15 lbs. The turbacil (2 mg/1 mL of 100 μL) was added for positive control and simple DMSO (Dimethyl sulfoxide) was added to test tube for negative control of fungus. Various concentrations of CuNPs (100 μg/150 μL, 500 μg/200 μL, 1000 μg/250 μL) were added to each strain of the test tube. The test tubes having SDA media were shaken and allowed to settle down overnight perpendicular to solidify the media in test tubes. Fungus of each strain was added to the top of the media of the test test-tube and incubated for seven days at 37 °C. After the incubation period, one can see the results as shown in Table 2.
3. Results and Discussion
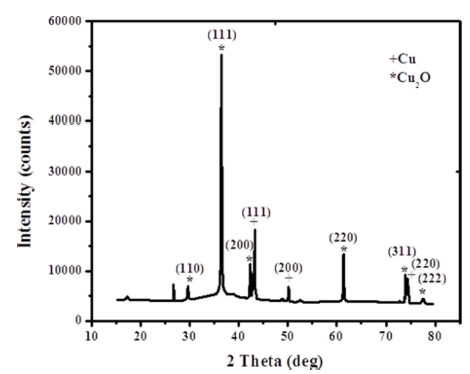
The XRD analysis established the crystal structure and size of the CuNPs. The x-ray diffraction pattern is shown in Fig. 2, wherein, the reflection bands observed at 2θ values of 43.51°, 50.46° and 74.17° correspond to the (111), (200) and (220) planes of metallic copper. Other bands located at 2θ values of 29.66°, 36.37°, 42.38°, 61.48°, 73.55° and 77.58° correspond to (110), (111), (200), (220), (311) and (222) planes of Cu2O, correspondingly, exhibit the shape of cubic Cu2O nano structures as described earlier in the literature.15,16) Thus, the X-ray profile shows that the resulted product is a combination of metallic Cu and Cu2O. Where, the mean crystalline size of copper and cuprous oxide nanoparticles was found out from the main diffraction peaks employing the Scherrer formula and were found to be of about 29.21 and 25.33 nm, respectively.
The XRD profile confirmed that both the crystalline phases of copper and cuprous oxide are coexisting. It reveals the formation of zerovalent copper nano structures in the process which is also simultaneously oxidized into Cu2O due to limited stability factors of Cu as described earlier.17,18) Thus, all the existing nanocrystals were found to be of Cu and Cu2O having no sign of cupric oxide (CuO). Similarly, the existence of nanocrystals could be confirmed from the peak widening in the X-ray pattern.19)
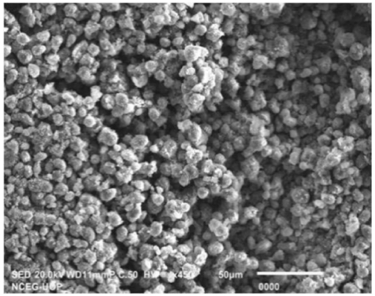
The SEM image of the CuNPs is provided in Fig. 3, Wherein, the image reveals that CuNPs are cubic in shape, having an agglomerated and aggregated form. During the synthesis process, the agglomeration may be occurring due to the high affinity of copper nuclei to bond or it may be happening due to the high surface area of the CuNPs as reported earlier in the literature.20,21)
In this chemical reduction method, metallic copper and cuprous oxide nanocrystals have been obtained from an aquous solutions of copper sulphate (as precursor), starch (as stabilizer), sodium hydroxide (as an alkaline medium) and an ascorbic acid (as reducing agent). Wherein, the ascorbic acid performs to develop elemental copper nanoparticles by reduction of copper ions disseminated in starch. The addition of NaOH executed the reduction rate. Along with the production of metallic copper, large amount of cuprous oxides are also found due to the oxidation of copper in the presence of small quantity of oxygen. Similarly, augmentation of NaOH is also anticipated to form copper (II) hydroxide (Cu (OH)2) which is afterwards found to produce Cu2O nanoparticles upon reduction reaction. The starch works as a dispersing agent to isolate copper ions from each other and as a consequence delivers improved size control of particles on nano scale. The presence of alkyl functional groups on the surface of metal cluster plays an important role to control the conversion of zerovalent copper to their oxides. As naturally, the formation of metal particles in liquid phase is unstable, and these particles tend to aggregate so as to reduce the total surface energy. Similarly, the van der Waals forces of attraction in between the metal crystals are also considered to be responsible for their agglomeration as reported earlier in the literature.13,17,21-23)
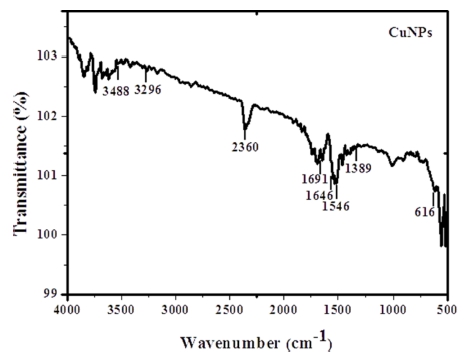
The profile of FTIR-ATR spectrum of CuNPs is provided in the Fig. 4. The numerous bands can be seen in the wave number region 3,296.03 cm-1, 1,691.87 cm-1, 1,646.21 cm-1, 1,546.20 cm-1 and 1,389.87 cm-1 corresponding to hydroxyl, scissor bending vibration mode of molecular H2O, acidic asymmetric stretching and C-H deformation of –CH2 or –CH3 aliphatic groups, respectively. 22) A band at wave number 2,360 cm-1 may be attributed to the atmospheric CO2. Another peak present at 616 cm-1 could be ascribed to the Cu-O vibrations. Further, the broad bands displayed at around 3,488 cm-1 and 616 cm-1 also may be assigned to the presence of OH surface stretching groups in CuNPs.11,23)
The engineered nanoparticles of Cu were tested for antibacterial activity against pathogenic bacterial strains. Maximum antibacterial activity against Staphylococcus aureus, Pseudomonas aeruginosa and Klebsiella pneumoniae was found at 200 μg/mL of the CuNPs as given in Table 1. It was worthy to note that the CuNPs were highly effective in the inhibition of growth of Gram -ve bacteria. The bacterial cells have extensive number of carboxylic groups in lipoproteins of cell membranes which upon release makes their outer surface as negatively charged.24) The presence of opposite charges on Cu2+ ions released from copper nanoparticles and bacterial cells causes their adhesion due to electrostatic forces. After binding with the bacterial cell the enterance of Cu2+ ions is increased inside the cytoplasm resulting in bioactivity of the copper nanoparticles.25) Moreover, copper nanoparticles kill bacteria by destrying cell wall and integrity of cell membrane, causing shrinkage of cellular vacuole and destruction of proteins.26)
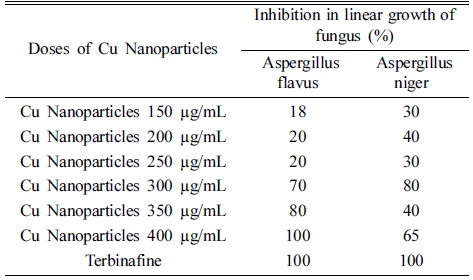
The antifungal activity of the Cu nanoparticles was concentrated dependant. Higher inhibition (%) in linear growth of the tested fungi viz. Aspergillus flavus and Aspergillus niger was recorded at 400 μg/mL of the Cu nanoparticles. Between the two tested fungal strains, Aspergillus flavus seems as more susceptible to the Cu nanoparticles as given in Table 2. Similar studies have reported antifungal properties of copper nanoparticles.27 Copper nanoparticles destroys integrity of cell membrane by causing lipid peroxidation, denaturation of proteins and nucleic acids and other cell biomolecules leading to cell death.28)
4. Conclusions
The present study demonstrated the synthesis and characterization of copper nanoparticles (CuNPs) through a simple and cost effective process of chemical reduction method, and their antibacterial/antifungal activities against the tested Gram-positive bacteria, and Gramm-negative bacteria. X-ray results designated the cubic shape of Cu and Cu2O nanocrystals having an average size 29.21 and 25.33 nm, respectively. The FTIR analysis confirmed the presence of functional groups. This low cost and simple fabrication process could be applied for mass production of Cu and Cu2O nanoparticles. The synthesized CuNPs also exhibited antimicrobial activity against the tested pathogenic bacterial and fungal strains, and thus have potential for utilization as drug carriers.